approaches only make use of t he topological structure
information from the vertices and fail to take into con-
sideration the roles of edges. It, however, is unreason-
able to ignore th e roles of edges, say the weak tie theory
[35] and percolation [36], since an edge may play an
important role in enhancing the locality or be significant
in maintaining the global connectivity. For example, the
famous weak ties theory indicates the job opport unities
and new ideas are usually from p ersons with weak con-
nections. Furthermore, the wea k ties can be used to
characterized the topological properties of networks
such as the stability of biological functions [37], the
accuracy of network structure prediction [38], the struc-
ture in mobile communication networks [39]. And the
percolation characterizes the tendency to undergo a
topological phase transition as the number of connec-
tions is progressively increased. Motivated by these
observations, we pose the following question:
Question: whet her the r oles of e dges can be used i n
protein complexes detection?
In this study, we aim to investigate the possibility to
extract protein complexes by exploring the roles of
edges and develop an affirmative answer to the above
question. In detail, similar to the weak ties effects in
mobile communication [39] and d ocument networks
[40], we prove complementa ry results on the PPI net-
works that is the edges connecting less similar nodes
are more significant in maintaining the global connectiv-
ity. By using the weak ties and percolation, a reliable
virtual network is constructed from the original PPI net-
work, in which each maximal clique corresponds to a
protein complex. A core-attachment based method is
developed. To test the performance of the proposed
algorithm, we applied it to the PPI networks. The
experimental results on the yeast PPI network show that
the proposed method outpe rforms DPClus [41], DEC-
AFF [42], MCL [14], MCODE [16] and Coach [24].
Further, the analysis of detected modules by the present
algorithm suggests that most of these modules have well
biologi cal significance in context of complexes, suggest-
ing that the roles of edges are critical in discovering
protein complexes.
Materials and methods
The key idea behind our algorithm consists of three
main steps: (1) verifying the existence of weak ties effect
in PPI networks; (2) constructing a rel iable network by
exploring the roles of edges; and (3) identifying the pro-
tein complexes by using a core-attachment based
method. We show them in turns.
Weak ties phenomenon in PPI networks
A network consists of two basic elements: vertices and
edges. Many measurements are developed to character-
ize the role of a node for structure and function includ-
ing random walk-based indices [43], PageRank score
[44]. In comparison, the study of the edge’ sroleisless
extensive.
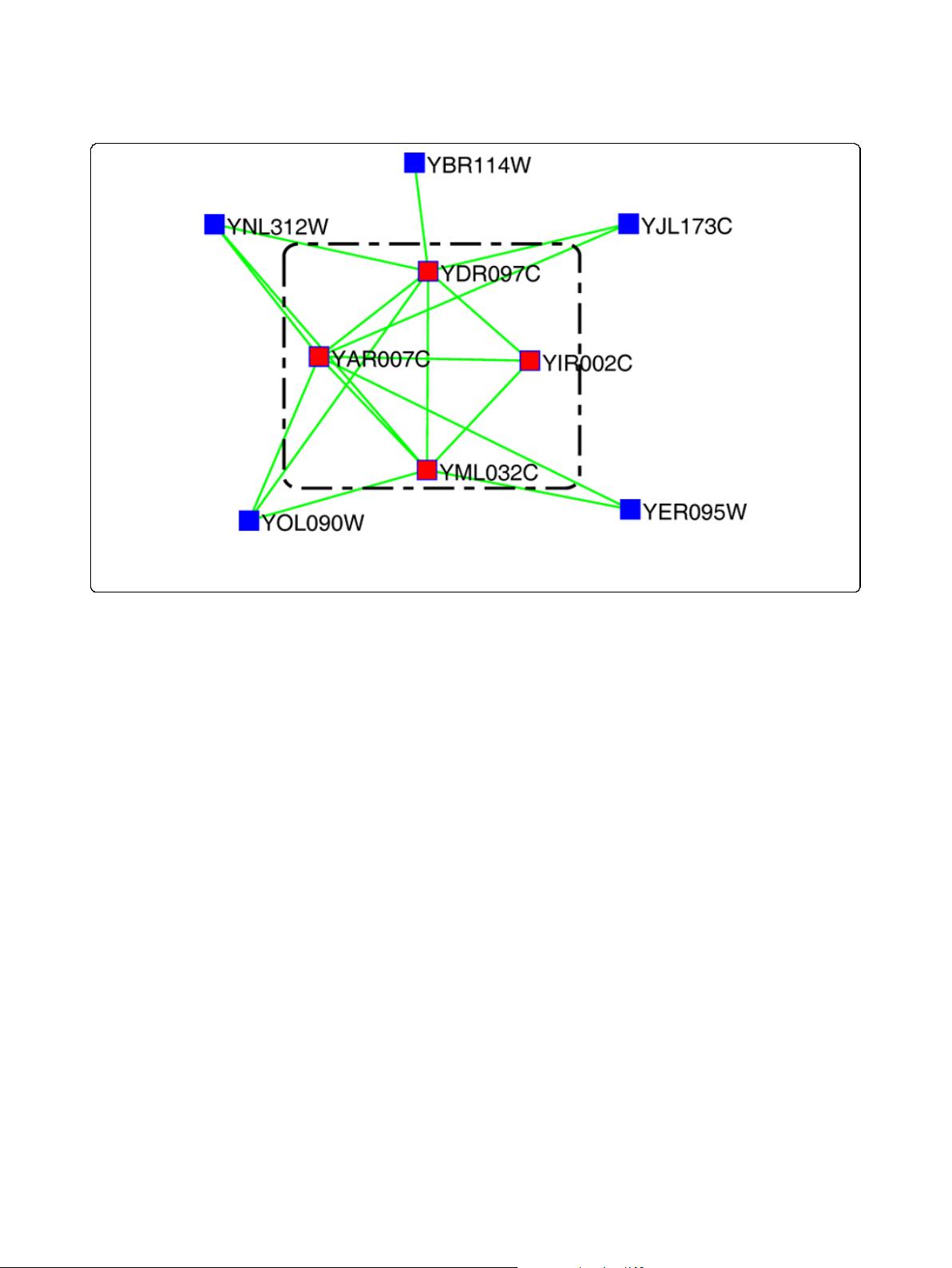
Figure 1 An schematic example of core-attachment structure of protein complexes. An exam ple o f the DNA repair complex [8], whose
core consists of four red proteins in the dotted square and others are the attachments of this complex. The interactions in this figure are from
the DIP data.
Ma and Gao BMC Systems Biology 2012, 6(Suppl 1):S6
http://www.biomedcentral.com/1752-0509/6/S1/S6
Page 3 of 15









