没有合适的资源?快使用搜索试试~ 我知道了~
首页深度学习驱动的医疗数据挖掘:从结构化到非结构化
深度学习驱动的医疗数据挖掘:从结构化到非结构化
需积分: 6 1 下载量 119 浏览量
更新于2024-07-09
收藏 32.27MB PDF 举报
“Advances in Mining Heterogeneous Healthcare Data” 是一篇关于在医疗健康领域利用深度学习技术挖掘异构数据的教程。这篇教程由 Fenglong Ma, Muchao Ye, Junyu Luo, Cao Xiao 和 Jimeng Sun 准备,涵盖了电子健康记录(EHR)的不同类型数据、各种应用以及深度学习在结构化和非结构化医疗数据挖掘中的最新进展。
电子健康记录(EHR)是患者在医疗机构多次就诊过程中生成的长期健康信息记录。从2008年至2015年,美国医院对电子健康记录系统的采用率显著增加,至2017年,94%的医院使用EHR数据进行临床实践指导的医院流程。EHR数据的应用范围广泛,包括但不限于疾病诊断、风险预测和治疗建议。
教程分为两大部分:
1. 挖掘结构化健康数据:
- 计算表型(Phenotyping):通过分析结构化的医疗数据,如病历、实验室结果和药物处方,识别并定义患者的临床表型,有助于研究疾病模式。
- 疾病检测/风险预测:使用深度学习模型来提前识别疾病迹象,提高早期诊断的准确性,同时预测疾病风险,从而实现预防性医疗。
- 治疗建议:根据患者的个人健康状况,利用深度学习算法生成个性化的治疗方案,优化治疗效果。
2. 挖掘非结构化健康数据:
- 自动化ICD编码:利用深度学习自动为临床记录分配国际疾病分类(ICD)代码,提高编码效率和准确性。
- 可理解的医学语言翻译:通过深度学习模型将医学术语转化为通俗易懂的语言,增强医患沟通。
- 医学报告生成:基于深度学习的自然语言处理技术,自动生成详细的医学检查报告,减轻医生的工作负担。
- 临床试验挖掘:在大量的医学文献中搜索和匹配适合的临床试验,加速新药或疗法的研发进程。
教程面向对将深度学习应用于医疗保健感兴趣的初级和高级学生、工程师和研究人员,对先验知识的要求较低。教程结束时会进行开放式问题讨论和问答环节,鼓励参与者深入探讨和互动。
总结来说,这篇教程旨在通过介绍最新的深度学习方法,推动医疗健康数据的高效利用,解决结构化和非结构化数据的挑战,促进医疗保健领域的科技进步。通过学习,参与者可以了解到如何利用深度学习技术来改善疾病诊断、提高医疗服务质量和效率,以及推动医学研究的发展。
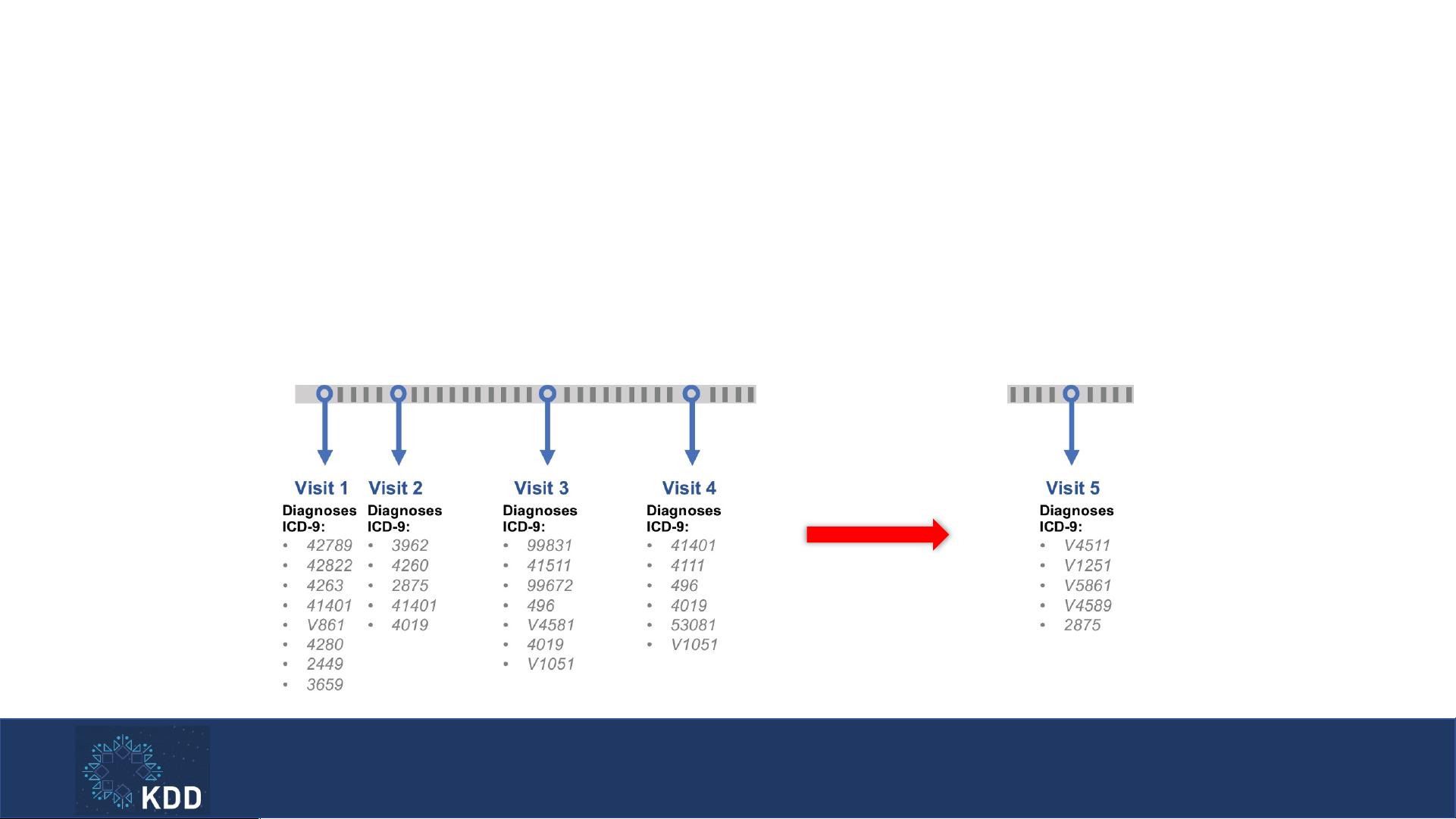
Phenotyping
• Goal: Learning medical concept representations from EHR data
• Approach: Predicting the next visit information according to all the
previous visits
16
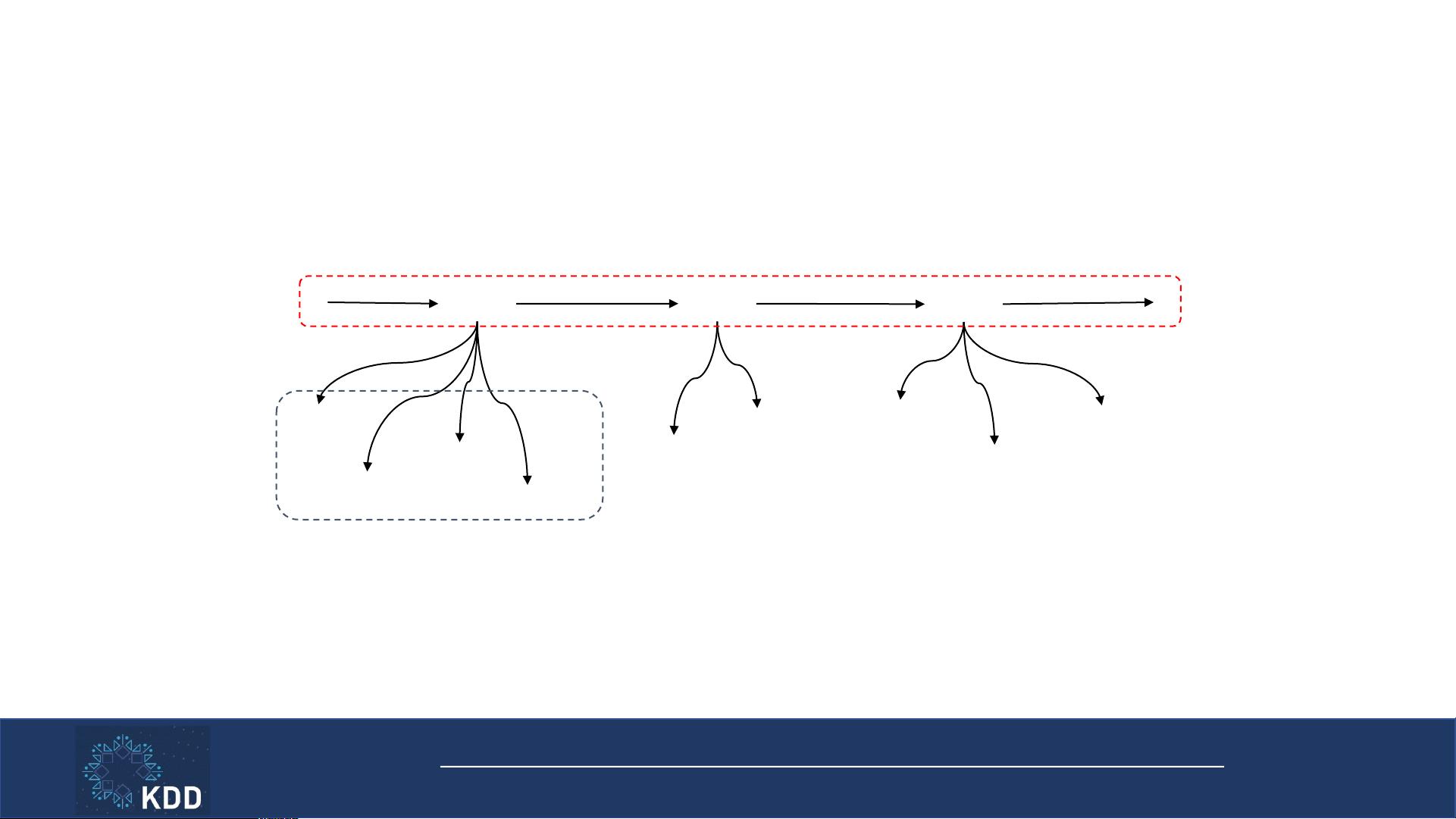
Med2Vec
• Two-layered representation learning
17
v Choi et al.
Med2Vec: Multi-layer Representation Learning for Medical Concepts
. KDD
2016.
Cough
Visit 1
Fever
Fever
Visit 2
Chill
Fever
Visit 3
Pneumonia
Chest X-ray
Tylenol
IV fluid
2. Sequential relation
1. Co-occurrence
• Objective function: the sum of
1. Negative intra-visit Skip-gram
• Because Skip-gram objective function is to be maximized
2. Inter-visit multi-label classification loss
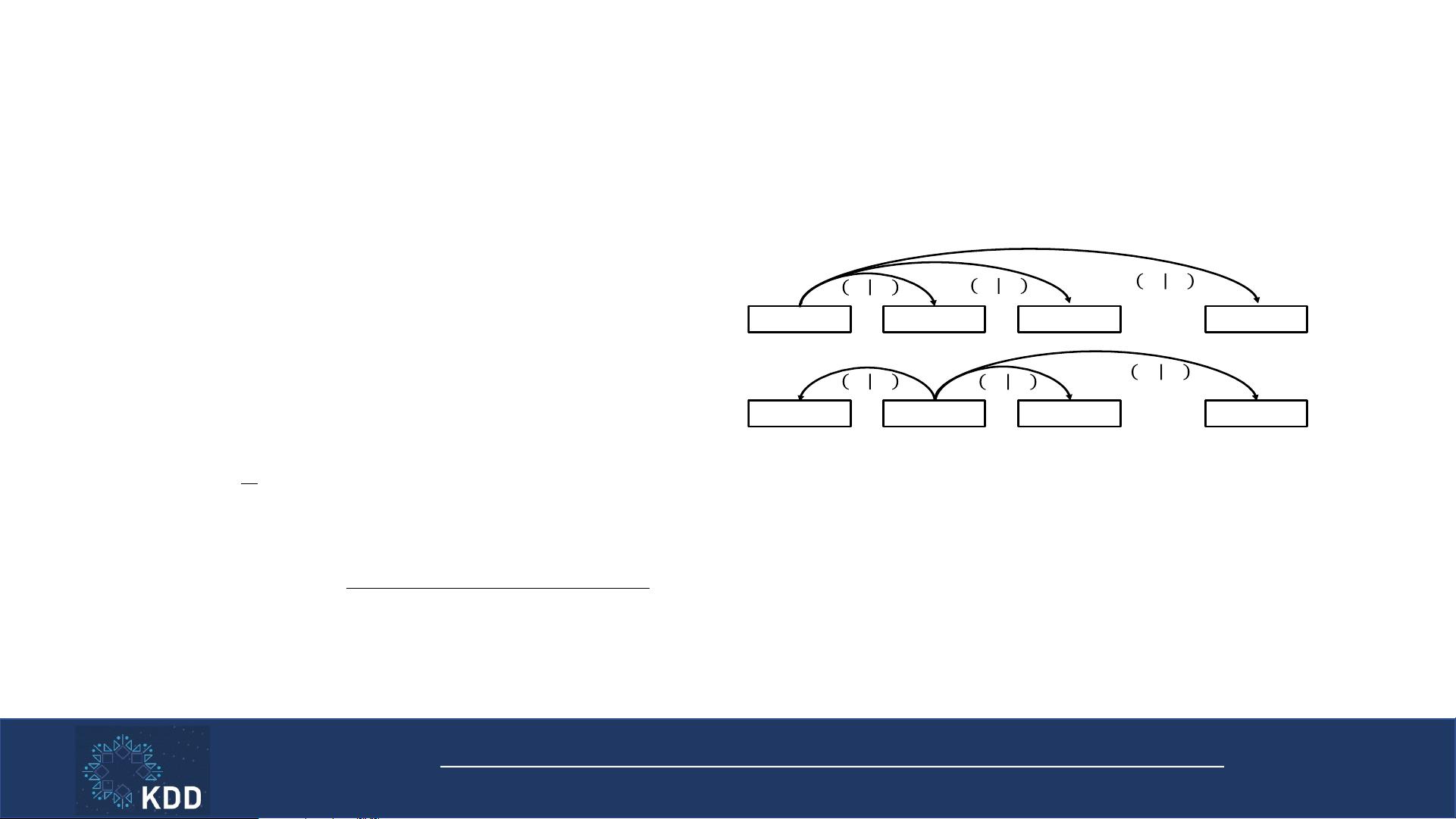
Intra-visit Skip-gram
• Model all pairs of medical codes in a visit
18
• Visit contains codes {c
1
, c
2
, c
3
, … c
n
}
• c
i
: i-th code among the code vocabulary C
• p(c
i
| c
j
): Skip-gram probability (see below)
• Each code c
1
, c
2
, c
3
, … c
n
is used as the ”input”
• Learn W
c
, the code representation
c
1
c
2
c
3
c
n
. . .
! "
#
"
$
! "
%
"
$
! "
&
"
$
c
1
c
2
c
3
c
n
. . .
! "
#
"
&
! "
$
"
&
! "
%
"
&
. . .
Repeat for n times
• V
t
: t-th visit
• c
i
, c
j
: codes in the visit V
t
• [:, j]: j-th column of the matrix
text window size, exp the element-wise exponential function,
and 1 denotes an all one vector. We have used MATLAB’s
notation for selecting a row in W
s
and a coordinate of b
s
.
3.3 Learning from the code-level information
As we described in the introduction, healthcare datasets
contain two-level information: visit-level sequence informa-
tion and code-level co-occurrence information. Since the loss
function in E q. (2)canefficiently capture the sequence level
information, now we need to find a way to use the second
source of information, i.e., the intra-visit co-occurrence of
the codes.
A natural choice to capture the code co-occurrence infor-
mation is to use Skip-gram. The main idea would be that
the representations for the codes that occur in the same visit
should predict each other. To embed Skip-gram in Med2Vec,
we can train W
c
2 R
m⇥|C|
(which also produces intermedi-
ate visit l evel representations) so that the i-th column of W
c
will be the representation for the i-th medical code among
total |C| co des. Note that given the unordered nature of
the codes inside a visit, unlike the original Skip-gram, we do
not distinguish between the “input” medical code and the
“output” medical code. In text, it is sensible to assume that
a word can serve a di↵erent role as a center word and a
context word, whereas in EHR datasets, we cannot classify
codes as center or context codes. It is also desirable to learn
the representations of di↵erent types of codes (e.g. di agno-
sis, medication, procedure code) in the same latent space so
that we can capture the hidden relationships between them.
However, precise interpretation of Skip-gram codes will be
difficult as W
c
will have positive and negative values. For in-
tuitive interpretation, we should learn code representations
with non-negative values. Note that in Eq.(1), if the binary
vector x
t
is a one-hot vector, then the intermediate visit rep-
resentation u
t
becomes a code representation. Therefore,
using the Skip-gram algorithm, we train the non-negative
weight ReLU(W
c
) instead of W
c
. This will not only use
the intra-visit co-occurrence information, but also guaran-
tee non-negative code representations. Moreover, ReLU pro-
duces sparse code representations, which further facilitates
easier interpretation of the codes.
The code representations to b e learned is denoted as a
matrix W
0
c
= ReLU(W
c
) 2 R
m⇥|C|
. From a sequence of
visits V
1
,V
2
,...,V
T
, the code-level representations can be
learned by maximizing the following log-likelihood,
min
W
0
c
1
T
T
X
t=1
X
i:c
i
2V
t
X
j:c
j
2V
t
,j6=i
log p(c
j
|c
i
), (3)
where p(c
j
|c
i
)=
exp
⇣
W
0
c
[:,j]
>
W
0
c
[:,i]
⌘
P
|C|
k=1
exp
⇣
W
0
c
[:,k]
>
W
0
c
[:,i]
⌘
. (4)
3.4 Unified training
The single unified framework can be obtained by adding
the two objective functions (3) and (2) as follows,
argmin
W ,b
1
T
T
X
t=1
n
X
i:c
i
2V
t
X
j:c
j
2V
t
,j6=i
log p(c
j
|c
i
)
+
X
w kw,k6=0
x
>
t+k
log
ˆ
y
t
(1 x
t+k
)
>
log(1
ˆ
y
t
)
o
By combining the two objective functions we learn both
code representations and visit representations from the same
source of patient visit records, exploiting both intra-visit
co-occurrence information as well as inter-visit sequential
information at the same time.
3.5 Interpretation of learned representations
While the original Skip-gram learns code representations
that have interesting properties such as additivity, in health-
care we need stronger interpretability. We need to be able
to associate clinical meaning to each dimension of both code
and visit representations. Interpreting the learned represen-
tations is based on analyzing each coordinate in both code
and visit embedding spaces.
Interpreting code representations.
If information is properly embedded into a lower dimen-
sional non-negative space, each coordinate of the lower di-
mension can be readily interpreted. Non-negative matrix
factorization (NMF) is a go od example. Since we trained
ReLU(W
c
) 2 R
m⇥|C|
, a non-negative matrix, to represent
the medical codes, we can employ a simple method to inter-
pret the meaning of each coordinate of the m-dimensional
code embedding space. We can find the top k codes that
have the largest values for the i-th coordinate of the code
embedding space as follows,
argsort(W
c
[i, :])[1 : k]
where argsort returns the indices of a vector that index its
values in a descending order. By studying the returned med-
ical codes, we can view each coordinate as a disease group.
Detailed examples are given in section 5.1
Interpreting visit representations.
To interpret the learned visit vectors, we can use the same
principle we used for interpreting the code representation.
For the i-th coordinate of the n-dimensional visit embed-
ding space, we can find the top k coordinates of the code
embedding space that have the strongest values as follows,
argsort(W
v
[i, :])[1 : k]
where we use the same argsort as before. Once we ob-
tain a set of code coordinates, we can use the knowledge
learned from interpreting the code representations to under-
stand how each visit coordinate is associated with a group
of diseases. This simple interpretation is possible because
the intermediate visit representation u
t
is a non-negative
vector, due to the ReLU activation function.
In the experiments, we also tried to find the input vector
x
t
that most activates the target visit coordinate [14, 21].
However, the results were very sensitive to the initial value of
x
t
, and even averaging over multiple samples were producing
unreliable results.
3.6 Complexity analysis
We first analyze the computational complexity of the code-
level objective function Eq. (3). Without loss of generality,
we assume the visit records of all patients are concatenated
into a single sequence of visits. Then the complexity for Eq.
(3) is as follows,
O(T M
2
|C|m)
text window size, exp the el ement-wise exponential function,
and 1 denotes an all one vector. We have used MATLAB’s
notation for selecting a row in W
s
and a coordinate of b
s
.
3.3 Learning from the code-level information
As we described in the introduction, healthcare datasets
contain two-level information: visit-level sequence informa-
tion and code-level co-occurrence information. Since the loss
function in Eq. (2)canefficiently capture the sequence level
information, now we need to find a way to use t he second
source of information, i.e., the intra-visit co-occurrence of
the codes.
A natural choice to capture the code co-occurrence infor-
mation is to use Skip-gram. The main idea would be that
the representations for the codes that occur in the same visit
should predict each other. To embed Skip-gram in Med2Vec,
we can train W
c
2 R
m⇥|C|
(which also produces intermedi-
ate visit level representations) so that the i-th column of W
c
will be the representation for the i-th medical code among
total |C| codes. Note that given the unordered nature of
the codes inside a visit, unlike the original Skip-gram, we do
not distinguish between the “input” medical code and the
“output” medical c ode. In text, it is sensible to assume that
a word can serve a di↵erent role as a center word and a
context word, whereas in EHR datasets, we cannot classify
codes as center or context codes. It is also desirable to learn
the representations of di↵erent types of codes (e.g. diagno-
sis, medication, procedure code) in the same latent space so
that we can capture the hidden relationships between them.
However, precise interpretation of Skip-gram codes will be
difficult as W
c
will have positive and negative values. For in-
tuitive interpretation, we should learn code representations
with non-negative values. Note that in Eq.(1), if the binary
vector x
t
is a one-hot vector, then the intermediate visit rep-
resentation u
t
becomes a code representation. Therefore,
using the Skip-gram algorithm, we train the non-negative
weight ReLU(W
c
) instead of W
c
. This will not only use
the intra-visit co-occurrence information, but also guaran-
tee non-negative code representations. Moreover, ReLU pro-
duces sparse code representations, which further facilitates
easier interpretation of the codes.
The code representations to be learned is denoted as a
matrix W
0
c
= ReLU(W
c
) 2 R
m⇥|C|
. From a sequence of
visits V
1
,V
2
,...,V
T
, the code-level representations can be
learned by maximizing the following log-likelihood,
min
W
0
c
1
T
T
X
t=1
X
i:c
i
2V
t
X
j:c
j
2V
t
,j6=i
log p(c
j
|c
i
), (3)
where p(c
j
|c
i
)=
exp
⇣
W
0
c
[:,j]
>
W
0
c
[:,i]
⌘
P
|C|
k=1
exp
⇣
W
0
c
[:,k]
>
W
0
c
[:,i]
⌘
. (4)
3.4 Unified training
The single unified framework can be obtained by adding
the two objective functions (3) and (2) as follows,
argmin
W ,b
1
T
T
X
t=1
n
X
i:c
i
2V
t
X
j:c
j
2V
t
,j6=i
log p(c
j
|c
i
)
+
X
w kw,k6=0
x
>
t+k
log
ˆ
y
t
(1 x
t+k
)
>
log(1
ˆ
y
t
)
o
By combining the two objective functions we learn both
code representations and visit representations from the same
source of patient visit records, exploiting both intra-visit
co-occurrence information as well as inter-visit sequential
information at the same time.
3.5 Interpretation of learned representations
While the original Skip-gram learns code representations
that have interesting properties such as additivity, in health-
care we need stronger interpretability. We need to be able
to associate clinical meaning to each dimension of both code
and visit representations. Interpreting the learned represen-
tations is based on analyzing each coordinate in both code
and visit embedding spaces.
Interpreting code representations.
If information is properly embedded into a lower dimen-
sional non-negative space, each coordinate of the lower di-
mension can be readily interpreted. Non-negative matrix
factorization (NMF) is a good example. Since we trained
ReLU(W
c
) 2 R
m⇥|C|
, a non-negative matrix, to represent
the medical codes, we can employ a simple method to inter-
pret the meaning of each coordinate of the m-dimensional
code embedding space. We can find the t op k co des that
have the largest values for the i-th coordinate of the code
embedding space as follows,
argsort(W
c
[i, :])[1 : k]
where argsort returns the indices of a vector that index its
values in a descending order. By studying the returned med-
ical codes, we can view each coordinate as a disease group.
Detailed examples are given in section 5.1
Interpreting visit representations.
To interpret the learned visit vectors, we can use the same
principle we used for interpreting the code representation.
For the i-th coordinate of the n-dimensional v isit embed-
ding space, we can find the top k coordinates of the code
embedding space that have t he strongest values as follows,
argsort(W
v
[i, :])[1 : k]
where we use the same argsort as before. Once we ob-
tain a set of code coordinates, we can use the knowledge
learned from interpreting the code representations to under-
stand how each visit coordinate is associated with a group
of diseases. This simple interpretation is possible because
the intermediate visit representation u
t
is a non-negative
vector, due to the ReLU activation function.
In the experiments, we also tried to find the input vector
x
t
that most activates the target visit coordinate [14, 21].
However, the results were very sensitive to the initial value of
x
t
, and even averaging over multiple samples were producing
unreliable results.
3.6 Complexity analysis
We first analyze the computational complexity of the code-
level objective function Eq. (3). Without loss of generality,
we as sume the visit records of all patients are concatenated
into a single sequence of visits. Then the c omplexity for Eq.
(3) is as follows,
O(T M
2
|C|m)
text window size, exp the element-wise exponential function,
and 1 denotes an all one vector. We have used MATLAB’s
notation for selecting a row in W
s
and a coordinate of b
s
.
3.3 Learning from the code-level information
As we described in the introduction, healthcare datasets
contain two-level information: visit-level sequence informa-
tion and code-level co-occurrence information. Since the loss
function in Eq. (2)canefficiently capture the sequence level
information, now we need to find a way to use the second
source of information, i.e., the intra-visit co-occurrence of
the codes.
A natural choice to capture the code co-occurrence infor-
mation is to use Skip-gram. The main idea would be that
the representations for the codes that occur in the same visit
should predict each other. To emb ed Skip-gram in Med2Vec,
we can train W
c
2 R
m⇥|C|
(which also produces intermedi-
ate visit level representations) so that the i-th column of W
c
will be the representation for the i-th m edical code among
total |C| codes. Note that given the unordered nature of
the codes inside a visit, unlike the original Skip-gram, we do
not distinguish between the “input” medical code and the
“output” medical code. In text, it is sensible to assume that
a word can serve a di↵erent role as a center word and a
context word, whereas in EHR datasets, we cannot classify
codes as center or context codes. It is also desirable to learn
the representations of di↵erent types of codes (e.g. diagno-
sis, medication, procedure code) in the same latent space so
that we can capture the hidden relationships between them.
However, precise interpretation of Skip-gram codes will be
difficult as W
c
will have positive and negative values. For in-
tuitive interpretation, we should learn code representations
with non-negative values. Note that in Eq.(1), if the binary
vector x
t
is a one-hot vector, then the intermediate visit rep-
resentation u
t
becomes a code representation. Therefore,
using the Skip-gram algorithm, we train the non-negative
weight ReLU(W
c
) instead of W
c
. This will not only use
the intra-visit co-occurrence information, but also guaran-
tee non-negative code representations. Moreover, ReLU pro-
duces sparse code representations, which further facilitates
easier interpretation of the codes.
The code representations to be learned is denoted as a
matrix W
0
c
= ReLU(W
c
) 2 R
m⇥|C|
. From a sequence of
visits V
1
,V
2
,...,V
T
, the code-level representations can be
learned by maximizing the following log-likelihood,
min
W
0
c
1
T
T
X
t=1
X
i:c
i
2V
t
X
j:c
j
2V
t
,j6=i
log p(c
j
|c
i
), (3)
where p(c
j
|c
i
)=
exp
⇣
W
0
c
[:,j]
>
W
0
c
[:,i]
⌘
P
|C|
k=1
exp
⇣
W
0
c
[:,k]
>
W
0
c
[:,i]
⌘
. (4)
3.4 Unified training
The single unified framework can be obtained by adding
the two objective functions (3) and (2) as follows,
argmin
W ,b
1
T
T
X
t=1
n
X
i:c
i
2V
t
X
j:c
j
2V
t
,j6=i
log p(c
j
|c
i
)
+
X
w kw,k6=0
x
>
t+k
log
ˆ
y
t
(1 x
t+k
)
>
log(1
ˆ
y
t
)
o
By combining the two objective functions we learn both
code representations and visit representations from the same
source of patient visit records, exploiting both intra-visit
co-occurrence information as well as inter-visit sequential
information at the same time.
3.5 Interpretation of learned representations
While the original Skip-gram learns code representations
that have interesting properties such as additivity, in health-
care we need stronger interpretability. We need to be able
to associate clinical meaning to each dimension of both code
and visit representations. Interpreting the learned represen-
tations is based on analyzing each coordinate in both code
and visit embedding spaces.
Interpreting code representations.
If information is properly embedded into a lower dimen-
sional non-negative space, each coordinate of the lower di-
mension can be readily interpreted. Non-negative matrix
factorization (NMF) is a good example. Since we trained
ReLU(W
c
) 2 R
m⇥|C|
, a non-negative matrix, to represent
the medical codes, we can employ a simple method to inter-
pret the meaning of each coordinate of the m-dimensional
code embedding space. We can find the top k codes that
have the largest values for the i-th coordinate of the code
embedding space as follows,
argsort(W
c
[i, :])[1 : k]
where argsort returns the indices of a vector that index its
values in a descending order. By studying the returned med-
ical codes, we can view each coordinate as a disease group.
Detailed examples are given in section 5.1
Interpreting visit representations.
To interpret the learned visit vectors, we can use the same
principle we used for interpreting the code representation.
For the i-th coordinate of the n-dimensional visit embed-
ding space, we can find the top k coordinates of the code
embedding space that have the strongest values as follows,
argsort(W
v
[i, :])[1 : k]
where we use the same argsort as before. Once we ob-
tain a set of code coordinates, we can use the knowledge
learned from interpreting the code representations to under-
stand how each visit coordinate is associated with a group
of diseases. This simple interpretation is possible because
the intermediate visit representation u
t
is a non-negative
vector, due to the ReLU activation function.
In the experiments, we also tried to find the input vector
x
t
that most activates the target visit coordinate [14, 21].
However, the results were very sensitive to t he initial value of
x
t
, and even averaging over multiple samples were producing
unreliable results.
3.6 Complexity analysis
We first analyze the computational complexity of the code-
level objective function Eq. (3). Without loss of generality,
we assume the visit records of all patients are concatenated
into a single sequence of visits. Then the complexity for Eq.
(3) is as follows,
O(T M
2
|C|m)
text window size, exp the element-wise exponential function,
and 1 denotes an all one vector. We have used MATLAB’s
notation for selecting a row in W
s
and a coordinate of b
s
.
3.3 Learning from the code-level information
As we described in the introduction, healthcare datasets
contain two-level information: visit-level sequence informa-
tion and code-level co- occurrence information. Since the loss
function in E q. (2)canefficiently capture the sequence level
information, now we need to find a way to use the second
source of information, i.e., the intra-visit co-occurrence of
the codes.
A natural choice to capture the code co-occurrence infor-
mation is to use Skip-gram. The main idea would be that
the representations for the codes that occur in the same visit
should predict each other. To embed Skip-gram in Med2Vec,
we can train W
c
2 R
m⇥|C|
(which also produces intermedi-
ate visit le vel representations) so that the i-th column of W
c
will be the representation for the i-th medical code among
total |C| codes . Note that given the unordered nature of
the codes inside a visit, unlike the original Skip-gram, we do
not distinguish betwe en the “input” m edical code and the
“output” medical code. In text, it is sensible to assume that
a word can serve a di↵erent role as a center word and a
context word, whereas in EHR datasets, we cannot classify
codes as center or context codes. It is also desirable to learn
the representations of di↵erent types of codes (e.g. diagno-
sis, medication, procedure code) in t he same latent space so
that we can capture the hidden relationships between them.
However, precise interpretation of Skip-gram codes will be
difficult as W
c
will have positive and negative values. For in-
tuitive interpretation, we should learn code representations
with non-negative values. Note that in Eq.(1), if the binary
vector x
t
is a one-hot vector, then the intermediate visit rep-
resentation u
t
becomes a code representation. Therefore,
using the Skip-gram algorithm, we train the non-negative
weight ReLU(W
c
) instead of W
c
. This will not only use
the intra-visit co-occurrence information, but also guaran-
tee non-negative code representations. Moreover, ReLU pro-
duces sparse code representations, which further facilitates
easier interpretation of t he codes.
The code representations to b e learned is denoted as a
matrix W
0
c
= ReLU(W
c
) 2 R
m⇥|C|
. From a sequence of
visits V
1
,V
2
,...,V
T
, the code-level representations can be
learned by maximizing the following log-likelihood,
min
W
0
c
1
T
T
X
t=1
X
i:c
i
2V
t
X
j:c
j
2V
t
,j6=i
log p(c
j
|c
i
), (3)
where p(c
j
|c
i
)=
exp
⇣
W
0
c
[:,j]
>
W
0
c
[:,i]
⌘
P
|C|
k=1
exp
⇣
W
0
c
[:,k]
>
W
0
c
[:,i]
⌘
. (4)
3.4 Unified training
The single unified framework can be obtained by adding
the two objective functions (3) and (2) as follows,
argmin
W ,b
1
T
T
X
t=1
n
X
i:c
i
2V
t
X
j:c
j
2V
t
,j6=i
log p(c
j
|c
i
)
+
X
w kw,k6=0
x
>
t+k
log
ˆ
y
t
(1 x
t+k
)
>
log(1
ˆ
y
t
)
o
By combining the two objective functions we learn both
code representations and visit representations from the same
source of patient visit records, exploiting both intra-visit
co-occurrence information as well as inter-visit sequential
information at the same time.
3.5 Interpretation of learned representations
While the original Skip-gram learns code representations
that have interesting properties such as additivity, in health-
care we need stronger interpretability. We need to be able
to associate clinical meaning to each dimension of both code
and visit representations. Interpreting the learned represen-
tations is based on analyzing each coordinate in both code
and visit embedding spaces.
Interpreting code representations.
If information is properly embedded into a lower dimen-
sional non-negative space, each coordinate of the lower di-
mension can be readily interpreted. Non-negative matrix
factorization (NMF) is a good example. Since we trained
ReLU(W
c
) 2 R
m⇥|C|
, a non-negative matrix, to represent
the medical codes, we can employ a simple method to inter-
pret the meaning of each coordinate of the m-dimensional
code embedding space. We can find the top k codes that
have the largest values for the i-th coordinate of the code
embedding space as follows,
argsort(W
c
[i, :])[1 : k]
where argsort returns the indices of a vector that index its
values in a descending order. By studying the returned med-
ical codes, we can view each coordinate as a disease group.
Detailed examples are given in section 5.1
Interpreting visit representations.
To interpret the learned visit vectors, we c an use the same
principle we used for interpreting the code representation.
For the i-th coordinate of the n-dimensional visit embed-
ding space, we can find the t op k coordinates of the code
embedding space that have the strongest values as follows,
argsort(W
v
[i, :])[1 : k]
where we use the same argsort as before. Once we ob-
tain a set of code coordinates, we can use the knowledge
learned from interpreting the code representations to under-
stand how each visit coordinate is associated with a group
of diseases. This simple interpretation is possible because
the intermediate visit representation u
t
is a non-negative
vector, due to the ReLU activation function.
In the experiments, we also tried to find the input vector
x
t
that most activates the target v isit coordinate [14, 21].
However, the results were very sensitive to the initial value of
x
t
, and even averaging over multiple samples were producing
unreliable results.
3.6 Complexity analysis
We first analyze the computational complexity of the code-
level objective function Eq. (3) . Without loss of generality,
we assume the visit records of all patients are concatenated
into a single sequence of visits. Then the complexity for Eq.
(3) is as follows,
O(T M
2
|C|m)
where
the representations for the codes that occur in the same v isit
should predict each other. To embed Skip-gram in Med2Vec,
we can train W
c
2 R
m⇥|C|
(which also produc es intermedi-
ate visit level representations) so that the i-th column of W
c
will be the representation for the i-th medical code among
total |C| codes. Note that given the unordered nature of
the codes inside a visit, unlike the original Skip-gram, we do
not distinguish between the “input” medical code and the
“output” medical code. In text, it is sensible to assume that
a word can serve a di↵erent role as a center word and a
context word, whereas in EHR datasets, we cannot classify
codes as center or context codes. It is also desirable to learn
the representat ions of di↵erent types of codes (e.g. diagno-
sis, medication, procedure code) i n the same latent space so
that we can capture the hidden relationships between them.
However, coordinate-wise interpretation of Skip-gram codes
is not straightforward because the positive and negative val-
ues of W
c
make it hard for each coordinate to focus on
a single coherent medical concept. For i ntuitive interpreta-
tion, we should learn code representations with non-negative
values. Note that i n Eq.(1), if the binary vector x
t
is a one-
hot vector, then the intermediate visit representation u
t
be-
comes a code representation. Therefore, using the Skip-gram
algorithm, we train the non-negative weight ReLU(W
c
) in-
stead of W
c
. This will not onl y use the intra-visit co-
occurrence i nformation, but also guarantee non-negative code
representations. Moreover, ReLU produces sparse code rep-
resentations, which further facilitates easier interpretation
of the codes.
The code representa tions to be learned is denoted as a
matrix W
0
c
= ReLU(W
c
) 2 R
m⇥|C|
. From a sequence of
visits V
1
,V
2
,...,V
T
, the code-level representations can be
learned by maximizing the following log-likelihood,
max
W
0
c
1
T
T
X
t=1
X
i:c
i
2V
t
X
j:c
j
2V
t
,j6=i
log p(c
j
|c
i
), (3)
where p(c
j
|c
i
)=
exp
⇣
W
0
c
[:,j]
>
W
0
c
[:,i]
⌘
P
|C|
k=1
exp
⇣
W
0
c
[:,k]
>
W
0
c
[:,i]
⌘
. (4)
3.4 Unified training
The single unified framework can be obtained by adding
the two objective functions (3) and (2) as follows,
argmin
W
c,v,s
,b
c,v,s
1
T
T
X
t=1
n
X
i:c
i
2V
t
X
j:c
j
2V
t
,j6=i
log p(c
j
|c
i
)
+
X
wkw,k6=0
x
>
t+k
log
ˆ
y
t
(1 x
t+k
)
>
log( 1
ˆ
y
t
)
o
By combining the two objective functions we learn both
code represent ations and visit representations from the same
source of patient visit records, exploiting both intra-visit
co-occurrence information as well as inter-visit sequential
information at the same time.
3.5 Interpretation of learned representations
While the original Skip-gram learns code representations
that have interesting properties s uch as additivity, in health-
care we need stronger interpretability. We need to be able
to associate clinical meaning to each dimension of both code
and visit representations. Interpreting the learned represen-
tations is based on analyzing each coordinate in both code
and visit embedding spaces.
Interpreting code representations.
If information is properly embedded into a lower dimen-
sional non-negative space, each coordinate of the lower di-
mension can be readily interpreted. Non-negative matrix
factorization (NMF) is a good example. Since we trained
ReLU(W
c
) 2 R
m⇥|C|
, a non-negative matrix, to represent
the medical codes, we can em ploy a simple method to inter-
pret the meaning of each coordi nate of the m-dimensional
code embedding space. We can find the top k codes that
have the largest values for the i-th coo rdinate of the code
embedding space as follows,
argsort(W
c
[i, :])[1 : k]
where argsort returns the indices of a vector that index its
values in a des cending order. By studying the returned med-
ical codes, we can view each coordinate as a disease group.
Detailed examples are given in section 5.1
Interpreting visit representations.
To interpret the learned visit vectors, we can use the same
principle we used for interpreting the code representation.
For the i-th coordi nate of the n-dimensional visit embed-
ding space, we can find the top k coordinates of the code
embedding space that have the s trongest values as follows,
argsort(W
v
[i, :])[1 : k]
where we use the same argsort as before. Once we ob-
tain a set of code coordinates, we can use the knowledge
learned from interpreting the code representations to under-
stand how each visit coordinate is associated with a group
of diseases. This simple interpretation is possible because
the intermediate visit representation u
t
is a non-negative
vector, due to the ReLU activation function.
In the experiments, we also tried to find the input vector
x
t
that most activates the target visit coordinate [14, 21].
However, the results were very sensitive to the initial value of
x
t
, and even averaging over multiple samples were producing
unreliable results.
3.6 Complexity analysis
We first analyze the computational complexity of the code-
level objective function Eq. (3). Without loss of generality,
we assume the visit records of all patient s are concatenated
into a single sequence of visits. Then the complexity for Eq.
(3) is as follows,
O(T M
2
|C| m)
where T is t he number of visits, M
2
is the average of squared
number of medical codes within a visit, |C| the number of
unique medical codes, m the size of the code representation.
The M
2
factor comes from iterating over all possible pairs
of codes within a visit. The complexity of the visit-level
objective function Eq.(2) is as follows,
O(Tw(|C|(m + n)+mn))
where w is the size of the context window, n the size of the
visit representation. The added terms come from generating
a visit representation via MLP. Since size of code represen-
tation m and size of visit representation n gene rally have the
v Choi et al.
Med2Vec: Multi-layer Representation Learning for Medical Concepts
. KDD
2016.
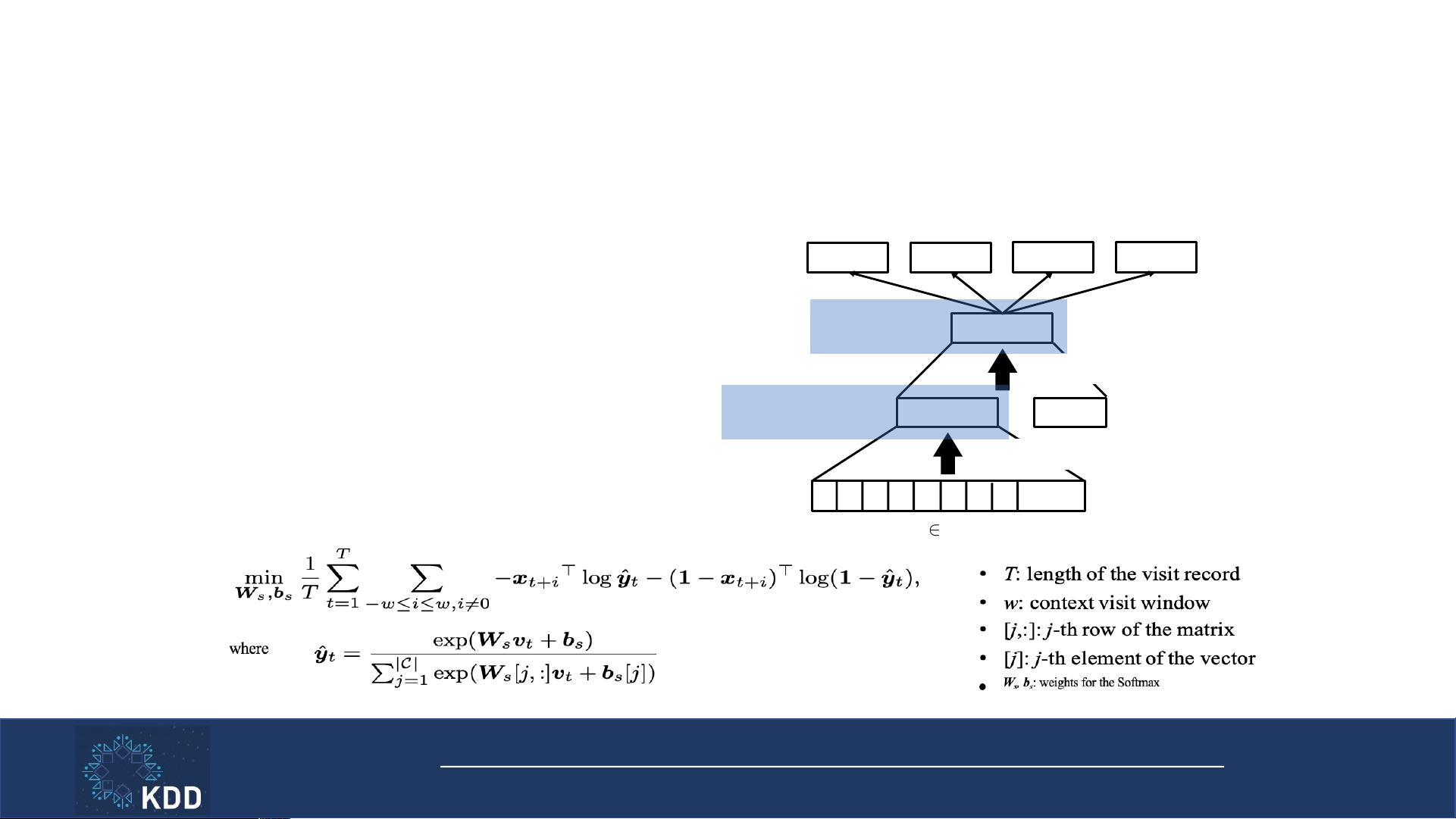
Inter-visit Multi-label Classification Loss
• Model relations between nearby visits
19
+
. . .
0
1 0 0 0 0 1 0
v
t
x
t+1
u
t
d
t
x
t-1
x
t
Softmax
x
t-2
x
t+2
. . . . . .
ReLU(W
v
[u
t
, d
t
] + b
v
)
ReLU(W
c
x
t
+ b
c
)
{0, 1}
|C|
Final visit
representation
• x
t
: one-hot coded Dx, Rx, Pr at time t
• u
t
: intermediate visit representation
• d
t
: patient demographic information
• v
t
: final visit representation
• W
c
, W
v
, b
c
, b
v
: weights to learn
• |C|: number of unique medical codes
Intermediate visit
representation
v Choi et al.
Med2Vec: Multi-layer Representation Learning for Medical Concepts
. KDD
2016.
Dipole
•
Imitate doctors’ diagnosis procedure + disease progression
20
Doctor Diagnosis
Disease Progression
l 558.9
l 477.9
l 401.9
l 274.9
l 530.8
l 278.0
l 584.9
l 995.91
l 518.81
l 786.50
l 564.00
l 357.0
l 305.02
l 852.20
l 959.09
l E849.7
l 723.1
l E888.9
l 959.01
l V49.84
l 300.00
l 305.02
l 530.81
l 786.50
l 401.9
l V58.61
l 786.50
l 428.0
l 780.2
Visit 1
Visit 2
Visit 3 Visit 4
Visit 5
Visit 6
Diagnoses
ICD-9 Codes:
Diagnoses
ICD-9 Codes:
Diagnoses
ICD-9 Codes:
Diagnoses
ICD-9 Codes:
Diagnoses
ICD-9 Codes:
Diagnoses
ICD-9 Codes:
Importance for
the prediction
v Ma et al.
Dipole: Diagnosis Prediction in Healthcare via Attention-based Bidirectional
Recurrent Neural Networks
. KDD 2017.
剩余168页未读,继续阅读
2011-11-25 上传
2022-05-28 上传
2009-07-09 上传
2021-03-05 上传
2019-07-28 上传
2009-07-16 上传
努力+努力=幸运
- 粉丝: 2
- 资源: 136
上传资源 快速赚钱
 我的内容管理
展开
我的内容管理
展开
 我的资源
快来上传第一个资源
我的资源
快来上传第一个资源
 我的收益 登录查看自己的收益
我的收益 登录查看自己的收益 我的积分
登录查看自己的积分
我的积分
登录查看自己的积分
 我的C币
登录后查看C币余额
我的C币
登录后查看C币余额
 我的收藏
我的收藏  我的下载
我的下载  下载帮助
下载帮助

最新资源
- ES管理利器:ES Head工具详解
- Layui前端UI框架压缩包:轻量级的Web界面构建利器
- WPF 字体布局问题解决方法与应用案例
- 响应式网页布局教程:CSS实现全平台适配
- Windows平台Elasticsearch 8.10.2版发布
- ICEY开源小程序:定时显示极限值提醒
- MATLAB条形图绘制指南:从入门到进阶技巧全解析
- WPF实现任务管理器进程分组逻辑教程解析
- C#编程实现显卡硬件信息的获取方法
- 前端世界核心-HTML+CSS+JS团队服务网页模板开发
- 精选SQL面试题大汇总
- Nacos Server 1.2.1在Linux系统的安装包介绍
- 易语言MySQL支持库3.0#0版全新升级与使用指南
- 快乐足球响应式网页模板:前端开发全技能秘籍
- OpenEuler4.19内核发布:国产操作系统的里程碑
- Boyue Zheng的LeetCode Python解答集
安全验证
文档复制为VIP权益,开通VIP直接复制
 信息提交成功
信息提交成功